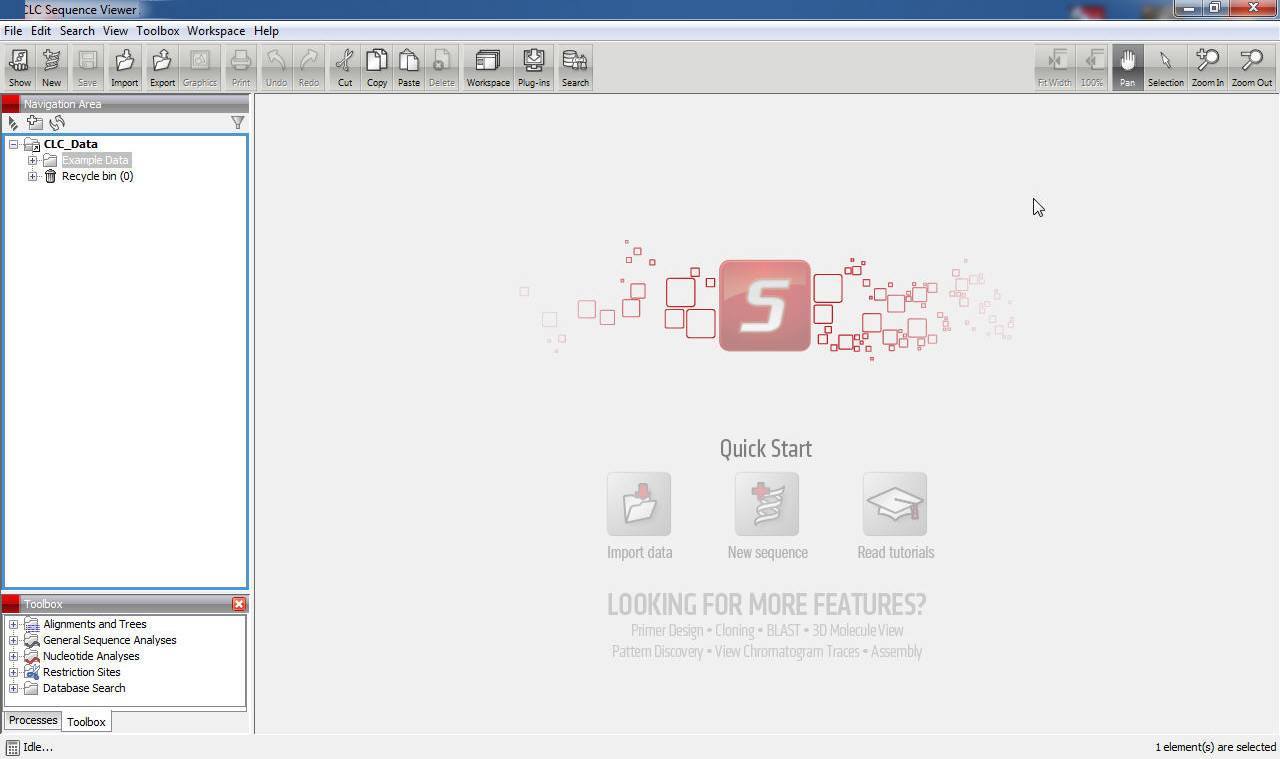
Then, you should save these genes for every animal you need (10at least ten), select saved genes and create new sequence list:Īnd then save in fasta format to obtain something like this:ĪTGACCAACCATCCCATCTTAATCAGCCTTATCATAGCCCTCTCCTACATCCTCCCCATTīut for further steps you might want to delete new lines characters, which can be easily done by using regular expression (notepad for Windows, gedit/vim for Linux): “$\n” (“$\r\n” for Windows) and then putting new lines before “>” signĬongratulations! Now you do have needed sequences in fasta format. Now you want to get on protein-coding gene and one ribosomal, so you should switch to circular view for the selected species: If you don’t have enough species (as in our case), you might want to search for “complete mitochondrion genome” or “mitochondrion”. We want to get order by accession, cause the best sequences are from NCBI and have NC_ prefix in front of them. So that’s what we put in the search box: “Passeriformes complete mitochondrion”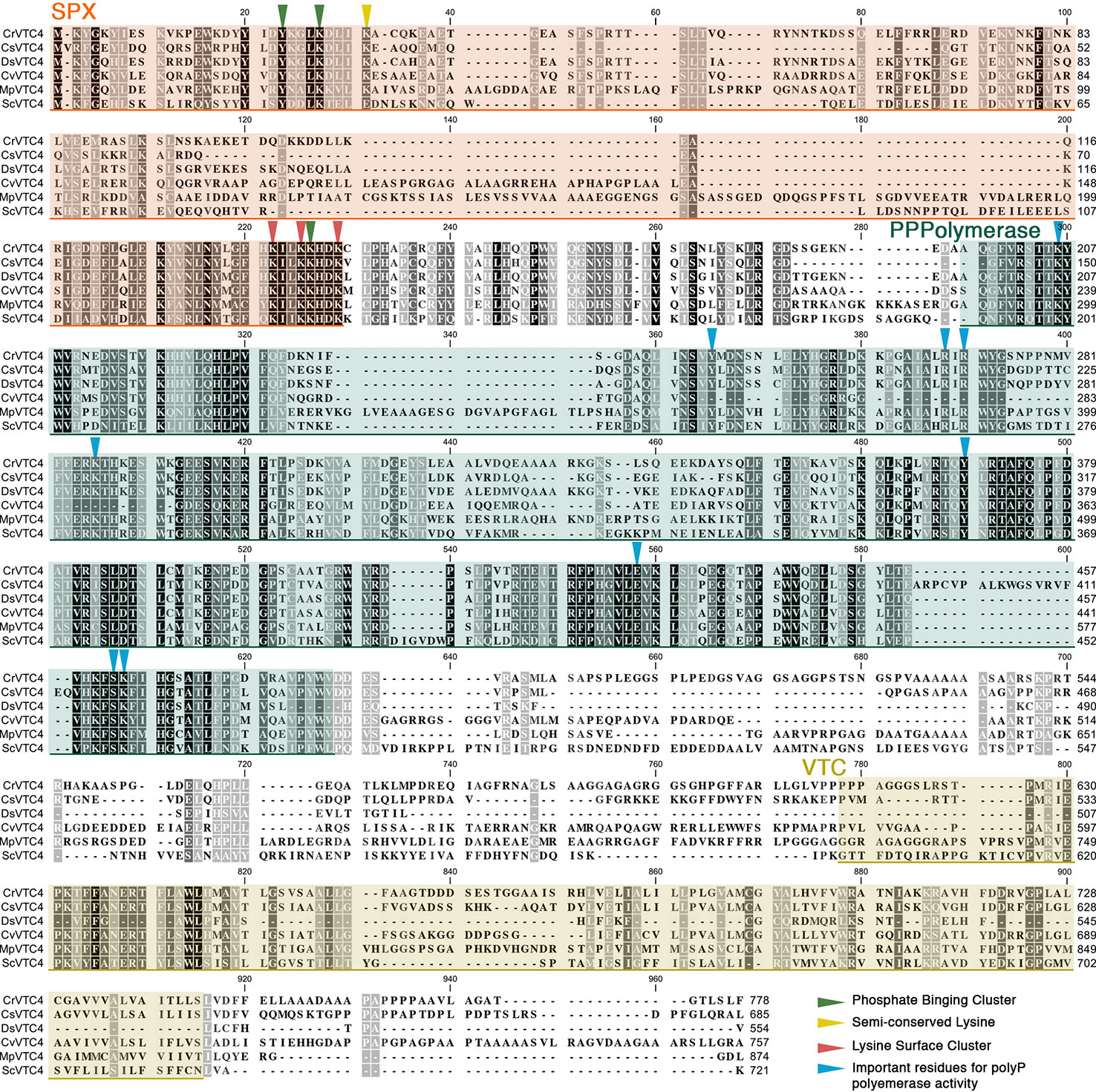 
Once you’ve chosen interesting animal/plant/whatever, you could go directly to wikipedia to find out its taxonomy, because if your target animal family doesn’t have much information, you’ll have to go one level above.ĭuring this tutorial we are going to explore Passeriformes, because the family Corvidae (crows) has too little information. This is one of the hardest parts, so this is should be done wisely, unless you are reading this, already having to build a particular tree, then it’s easy for you, cause you don’t have a choice. ALIGNMENT ALGORITHMS AND DATA MANAGINGĪt first we need to decide, for which group of animals we are going to build our phylogenetic tree. code nucleotide gaps as presence/absence data using GapCoder (which takes as input a modified FASTA file).ġ Execute ProtTest on the Phylip-formatted translated PCG dataset to identify the most suitable amminoacid substitution model Ģ use RAxML to estimate 100 maximum likelihood trees from the translated PCG dataset ģ create a 60%-threshold consensus tree using PhyUtility ġ Execute jModelTest on FASTA nucleotide dataset to identify the most suitable nucleotide substitution model Ģ concatenate the aminoacid and the nucleotide datasets (both data and presence/absence gap scores) in a single NEXUS file ģ load this NEXUS files with MrBayes and instruct the software for partitioning, modelling, and analyzing data ġ Use SplitsTree to perform different reconstructions of phylogenetic networks Ģ provide r8s the output tree of MrBayes to obtain an ultrametric tree (parameters will be optimized group by group). convert the translated PCG dataset into Phylip format using MEGA or CLC Sequence Viewer Ĩ. 
align the three datasets using M-Coffee, trying different combinations of algorithms ħ. translate the PCG dataset into amminoacids, using the correct mitochondrial code ĥ. extract the DNA sequences of a Protein Coding Gene (PCG) and a Ribosomal Gene (RG) and save the two datasets in FASTA format Ĥ. use CLC Sequence Viewer to download 15-20 complete mitochondrial genomes of the selected group ģ. Identify a zoological group of interest (aiming to reconstruct a deep phylogeny, this should be a higher-level taxon, like a Phylum, a Class, an Order, or similar) Ģ. SETTING UP THE DATASET: ALIGNMENT ALGORITHMS AND DATA MANAGINGġ. We are going to use the following workflow: Of course, there are more approaches available and you can use whatever you like / need to use, this is just an example, of how we can build a phylogenetic tree. This is going to be a « building a phylogenetic tree for dummies» manual. In this article we provide short manual for building a phylogenetic tree. What do we need for it? How is it done? What is it?įirst of all, phylogenetic tree is a branching diagram or “tree ” showing the inferred evolutionary relationships among various biological species or other entities-their phylogeny -based upon similarities and differences in their physical or genetic characteristics. It is the tool used by the biologists to see, how different population can be, what split is about to happen and how much is subjects of one population differ from other. Okay, we are about to build a phylogenetic tree. Twitter LinkedIn GitHub Value generation by Kirill | Bioinformatics How to build phylogenetic tree
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
